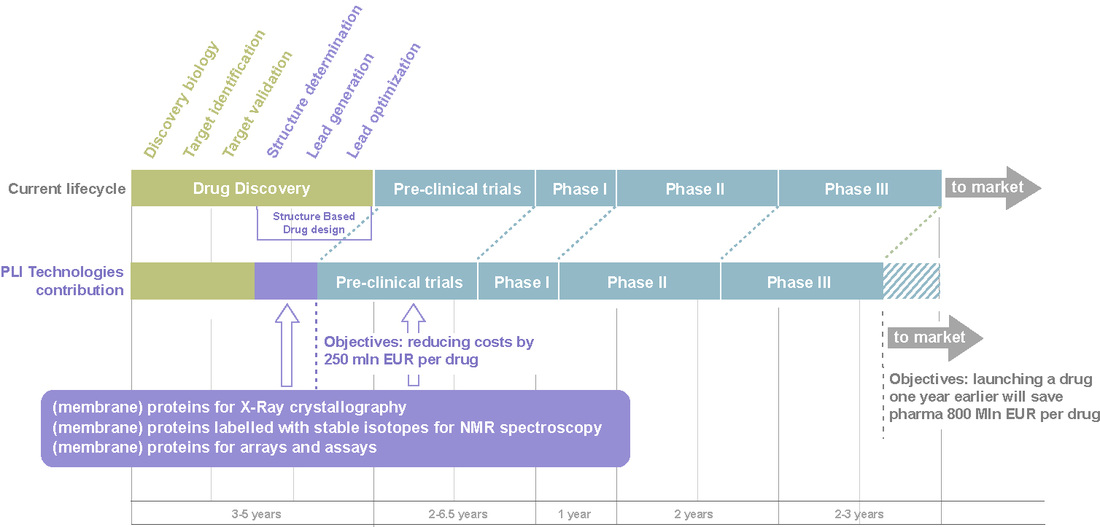
Speeding up drug design
An urgent need to develop new drugs
There is an urgent need to develop new drugs to fight old e.g. cancer, re- emerging e.g. tuberculosis and newly emerging e.g. AIDS, SARS diseases. The industry is becoming increasingly competitive and in order to be able to launch an increasing number of successful drugs each year, pharmaceutical companies must find faster and more efficient ways to discover compounds effective as drugs. Pharmaceutical companies therefore will have to expand their current drug discovery process, which involves testing usually more than 50,000 potential compounds to find a suitable drug compound (called “high throughput screening”) to a more targeted approach (“rational design”), that relies on a detailed knowledge of the protein structure. Once a 3D structure is known in sufficient detail and the essential interactive elements have been mapped, in silico modeling can be applied to design pharmacologically suitable families of compounds with a certain desired, possibly even novel physiological activity.
Human Drug Targets
The currently available drugs act on about 500 targets, more than 90 % of which are proteins. Main drug targets for drug therapy are G protein coupled receptors (GPCRs) and GPCR-based drugs represent more than 50% of all currently marketed drugs; this is estimated to be about 10% of all GPCRs, the receptors with unknown ligands being denoted as “orphan” receptors. The X-ray structures have been solved for only six GPCRs. Due to the functional genomics and proteomics efforts, the human genome will ultimately reveal at least 1500 and possibly up to 10.000 novel potential protein drug targets. This provides new business opportunities.
Business opportunities
The classical technique for obtaining the 3D structure of biological macromolecules at great atomic detail is X-ray diffraction of highly ordered 3-D crystals. Although to date the structure of over 10.000 soluble proteins has been determined at a resolution better than 3 Å, membrane proteins lag far behind with c.a. 200 entries in the protein structure database and 6 GPCRs. An alternative approach for protein structure determination and the only one for the proteins that can not be crystallized is nuclear magnetic resonance (NMR) spectroscopy. A major advantage of NMR is in that it allows the study of proteins in a physiological environment. This technique can provide information about chemical as well as electronic structure, and is applicable in a fluid-state as well as a solid-state mode.
However, both X-ray and NMR require large quantities (milligram scale) of highly purified proteins. In addition, NMR requires incorporation of the NMR active nuclei (such as 13C and 15N) in the molecule of interest. A suitable heterologous expression system for eukaryotic membrane proteins (drug targets), allowing both large scale functional production and affordable stable-isotope labelling is a major challenge.
There is an urgent need to develop new drugs to fight old e.g. cancer, re- emerging e.g. tuberculosis and newly emerging e.g. AIDS, SARS diseases. The industry is becoming increasingly competitive and in order to be able to launch an increasing number of successful drugs each year, pharmaceutical companies must find faster and more efficient ways to discover compounds effective as drugs. Pharmaceutical companies therefore will have to expand their current drug discovery process, which involves testing usually more than 50,000 potential compounds to find a suitable drug compound (called “high throughput screening”) to a more targeted approach (“rational design”), that relies on a detailed knowledge of the protein structure. Once a 3D structure is known in sufficient detail and the essential interactive elements have been mapped, in silico modeling can be applied to design pharmacologically suitable families of compounds with a certain desired, possibly even novel physiological activity.
Human Drug Targets
The currently available drugs act on about 500 targets, more than 90 % of which are proteins. Main drug targets for drug therapy are G protein coupled receptors (GPCRs) and GPCR-based drugs represent more than 50% of all currently marketed drugs; this is estimated to be about 10% of all GPCRs, the receptors with unknown ligands being denoted as “orphan” receptors. The X-ray structures have been solved for only six GPCRs. Due to the functional genomics and proteomics efforts, the human genome will ultimately reveal at least 1500 and possibly up to 10.000 novel potential protein drug targets. This provides new business opportunities.
Business opportunities
The classical technique for obtaining the 3D structure of biological macromolecules at great atomic detail is X-ray diffraction of highly ordered 3-D crystals. Although to date the structure of over 10.000 soluble proteins has been determined at a resolution better than 3 Å, membrane proteins lag far behind with c.a. 200 entries in the protein structure database and 6 GPCRs. An alternative approach for protein structure determination and the only one for the proteins that can not be crystallized is nuclear magnetic resonance (NMR) spectroscopy. A major advantage of NMR is in that it allows the study of proteins in a physiological environment. This technique can provide information about chemical as well as electronic structure, and is applicable in a fluid-state as well as a solid-state mode.
However, both X-ray and NMR require large quantities (milligram scale) of highly purified proteins. In addition, NMR requires incorporation of the NMR active nuclei (such as 13C and 15N) in the molecule of interest. A suitable heterologous expression system for eukaryotic membrane proteins (drug targets), allowing both large scale functional production and affordable stable-isotope labelling is a major challenge.
We offer
> cost effective labelling of high-value biomolecules of different complexity for metabolomics, proteomics, structural biology
> milligram scale production of recombinant (membrane) proteins
> license on Compositions and methods for stable isotope labeling of biological compounds
We seek strategic partners for:
> milligram scale production of recombinant (membrane) proteins
> license on Compositions and methods for stable isotope labeling of biological compounds
We seek strategic partners for:
- Out licensing of proprietary technology to label biomolecules with stable isotopes
- Joint R&D to improve efficiency and to speed up drug discovery process via:
- 3-D drug (target) structure determination
- Lead generation
- Lead optimization
Integrative multidisciplinary approach
We are proposing multidisciplinary approach to resolve 3D structure of drug target complex that requires the following: a) large scale membrane protein (drug target) production, b) stable isotope labelling, c) development of ssNMR and FTIR methodology for measurements of drug-drug target interactions and structural analysis. We are approaching this along the following experimental lines A-E.
A. Functional large scale expression and purification.
B. Stable-isotope labelling of ligands and receptors.
C. FT-IR studies to probe secondary structure and ligand induced conformational changes.
D. ssNMR studies to probe ligand conformation and ligand-protein cross-talk.
E. Molecular modeling.
We are proposing multidisciplinary approach to resolve 3D structure of drug target complex that requires the following: a) large scale membrane protein (drug target) production, b) stable isotope labelling, c) development of ssNMR and FTIR methodology for measurements of drug-drug target interactions and structural analysis. We are approaching this along the following experimental lines A-E.
A. Functional large scale expression and purification.
B. Stable-isotope labelling of ligands and receptors.
C. FT-IR studies to probe secondary structure and ligand induced conformational changes.
D. ssNMR studies to probe ligand conformation and ligand-protein cross-talk.
E. Molecular modeling.